AI predicts the structure of all known proteins and opens a new universe for science
The DeepMind lab, owned by the same company as Google, has calculated the predicted shapes of 200 million molecules, some of them essential for understanding devastating diseases such as Alzheimer’s or Parkinson’s
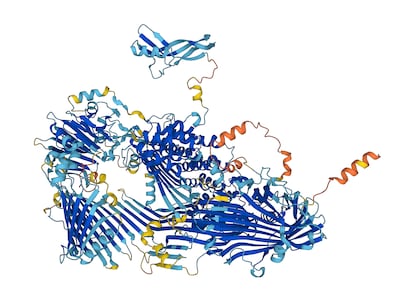
DeepMind’s artificial intelligence (AI) software has predicted the structure of nearly every known protein – about 200 million molecules. Knowing the structure of these molecules will help scientists understand the biology of every living thing on the planet, as well as how devastating diseases like malaria, Alzheimer’s and cancer develop.
“We’re at the beginning of new era of digital biology,” said Demis Hassabis, the AI and neuroscience expert who is the principal developer of AlphaFold, the neural network system that has almost completely solved one of the biggest challenges in the field of biology.
A child chess prodigy and expert video gamer, Hassabis is a British citizen who founded DeepMind in 2010, a company that creates artificial intelligence systems capable of learning like humans. In 2013, DeepMind developed a system that surpasses human level performance on Atari video games. The following year, Google announced that it had bought the company for US$500 million. In 2017, DeepMind’s AlphaGo system beat all the top players of Go, the highly complex Asian board game similar to chess. Hassabis then focused his company on a much bigger challenge – predicting the 3D shapes of proteins by reading their 2D gene sequences written in DNA letters.
Knowing the 3D structure of these molecules is essential for understanding how they function, but it is an immensely difficult problem to solve. Some have compared it with trying to put together a jigsaw puzzle with tens of thousands of blank pieces.
Without advanced technology, figuring out the structure or shape of a single protein composed of 100 basic units (amino acids) could take up to 13.7 billion years, the age of the universe. Some scientists using electron microscopy or huge particle accelerators such as the one at the European Synchrotron Radiation Facility in Grenoble (France) reduced the problem-solving time to several years. But Google’s AlphaFold system can determine the structure of a protein in just a few seconds.
“This protein universe is… a gift to humanity,” said Hassabis during a joint July 26 press briefing conference with the European Molecular Biology Laboratory (EMBL), an intergovernmental organization dedicated to molecular biology research that collaborated in AlphaFold’s development.
Before AlphaFold, it took 60 years and thousands of scientists to determine the structures of about 200,000 proteins. This research was used as learning material for AlphaFold, which searched for valid patterns that predict the shape of proteins. By 2021, it had successfully predicted the structures of a million proteins, including all human proteins. The latest release of AlphaFold results extends the number to 200 million proteins – virtually every known protein of every living thing on the planet.
DeepMind is providing free and open access to the AlphaFold code and protein database, both of which can be downloaded. A search of this “Google of life” database will display the 2D sequence of a protein and a 3D model with a corresponding level of reliability, which has a margin of error comparable to or lower than conventional prediction methods.
It is important to note that AlphaFold does not determine reality – it predicts reality. AlphaFold reads the genetic sequence and estimates the most likely configuration of its amino acids. The prediction has a high level of reliability, which saves a lot of time and money for scientists doing theoretical work, as they don’t need to use expensive equipment to determine the actual structure of a protein until absolutely necessary.
The applications of this new tool are virtually endless because microscopic proteins are involved in every conceivable biological process, such as bee colony collapse and crop heat resistance. A team led by Matt Higgins at the University of Oxford (UK) has used AlphaFold to help develop an antibody (a type of protein) that is capable of neutralizing one of the proteins that must be present for the malaria pathogen to reproduce. This could accelerate research to develop the first highly effective vaccine against the disease, thereby preventing mosquito transmission of the parasite.
More successes
Another AlphaFold-related success is the development of the most detailed nuclear pore structure available. Nuclear pores are a doughnut-shaped protein complex that is the gateway to the nucleus of human cells, and have been linked to a host of diseases, including cancer and cardiovascular disease. Jan Kosinski, an EMBL researcher and co-leader of the nuclear pore modeling effort, told EL PAÍS that AlphaFold provides scientists with unprecedented access to understanding how the recipe of life (written in the genome) works when translated into proteins.
Hassabis and his colleagues and DeepMind and EMBL say that they have analyzed the risks involved in making the AlphaFold system and data openly accessible. “The benefits clearly outweigh the risks,” said Hassabis, adding that it’s up to the international community to decide whether to restrict use of the technology as it develops further.
One of the most practical applications of AlphaFold is the design of tailor-made molecules that can block harmful proteins or, better yet, modulate their activity, a much more desirable effect when developing new drugs, said Carlos Fernández, a scientist with the Spanish National Research Council (CSIC) and leader of the structural biology group of the Spanish Society for Biochemistry and Molecular Biology (SEBBM). His team has used AlphaFold to predict part of the structure of a protein complex necessary for propagating the trypanosome found in sub-Saharan Africa that causes sleeping sickness.
Years of work now lie ahead to confirm the accuracy of AlphaFold’s predictions, says biologist José Márquez, an expert in protein structure at the European Synchrotron Radiation Facility in Grenoble. “The next frontier for AlphaFold will be its use in designing protein-blocking or protein-activating drugs, a problem they are already tackling,” said Márquez. And there’s another puzzle to solve: AlphaFold cannot say why a protein is shaped as it is, which could be an essential element of research on diseases like Alzheimer’s or Parkinson’s, both of which are related to misfolded proteins.
Alfonso Valencia, director of life sciences at the National Supercomputing Center in Barcelona (Spain), discusses some of the system’s shortcomings. “AlphaFold can’t solve everything because it can only predict what is in the domain of known things. For example, it cannot accurately predict the structure of proteins that protect against freezing because they are rare, and the databases don’t contain many samples. Nor can it predict the consequences of mutations, an issue of great concern to medicine,” said Valencia.
Valencia acknowledges the advantages of providing free and open access to AlphaFold, which enables other scientists to improve or modify the system as needed. “It’s clear that the DeepMind people are looking to win the Nobel Prize by acting transparently,” said Valencia. “It’s great for their image and gives them a competitive advantage over other companies like Facebook. On the other hand, they did hint that they might reserve specific health data for private use and drug development.”